 Find active subnetworks/pathway modules.For instance, users may select nodes involved in a threshold number of interactions, nodes that share a particular GO annotation, or nodes whose gene expression levels change significantly in one or more conditions according to p-values loaded with the gene expression data. Filter the network to select subsets of nodes and/or interactions based on the current data.Plugins are available for network and molecular profile analysis.Easily navigate large networks (100,000+ nodes and edges) by an efficient rendering engine.Use the bird's eye view to easily navigate large networks. 
And this structure can be saved in a session file. Use the network manager to easily organize multiple networks.Zoom in/out and pan for browsing the network.A variety of layout algorithms are available, including cyclic and spring-embedded layouts. according to user-configurable colors and visualization schemes. Expression data can be mapped to node color, label, border thickness, or border color, etc. View a superposition of gene expression ratios and p-values on the network.Customize network data display using powerful visual styles.Cytoscape session file includes networks, attributes (for node/edge/network), desktop states (selected/hidden nodes and edges, window sizes), properties, and visual styles. Load and save state of the cytoscape session in a cytoscape session (.cys) file.Directly import GO terms and annotations from OBO and Gene Association files.Import gene functional annotations from the Gene Ontology (GO) and KEGG databases.For example, input a set of custom annotation terms for your proteins, create a set of confidence values for your protein–protein interactions. Load and save arbitrary attributes on nodes and edges.Input mRNA expression profiles from tab- or space-delimited text files.Load and save networks and node/edge attributes in an XML document format called XGMML (eXtensible Graph Markup and Modeling Language).Load and save previously-constructed interaction networks in GML format (Graph Modelling Language). 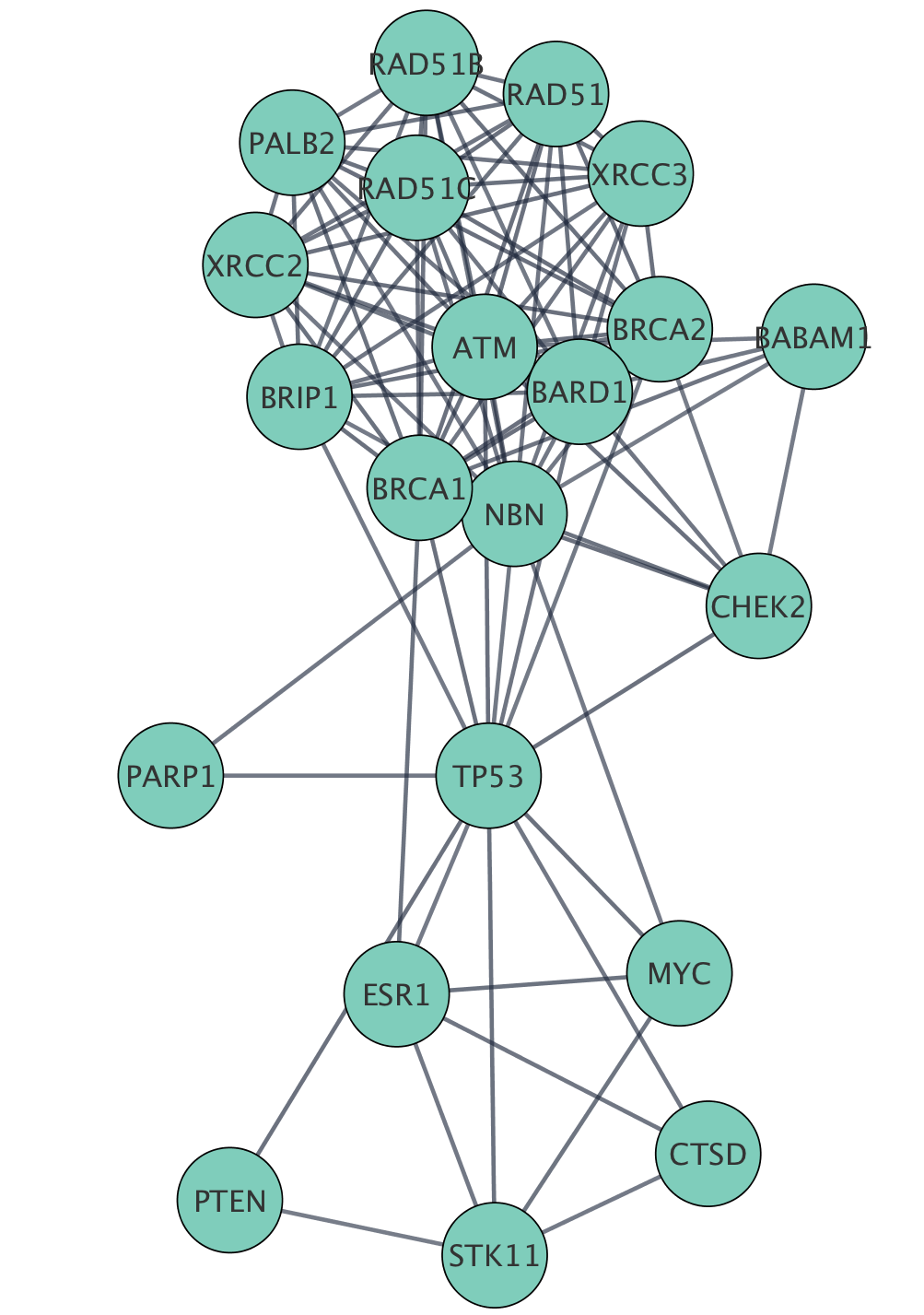 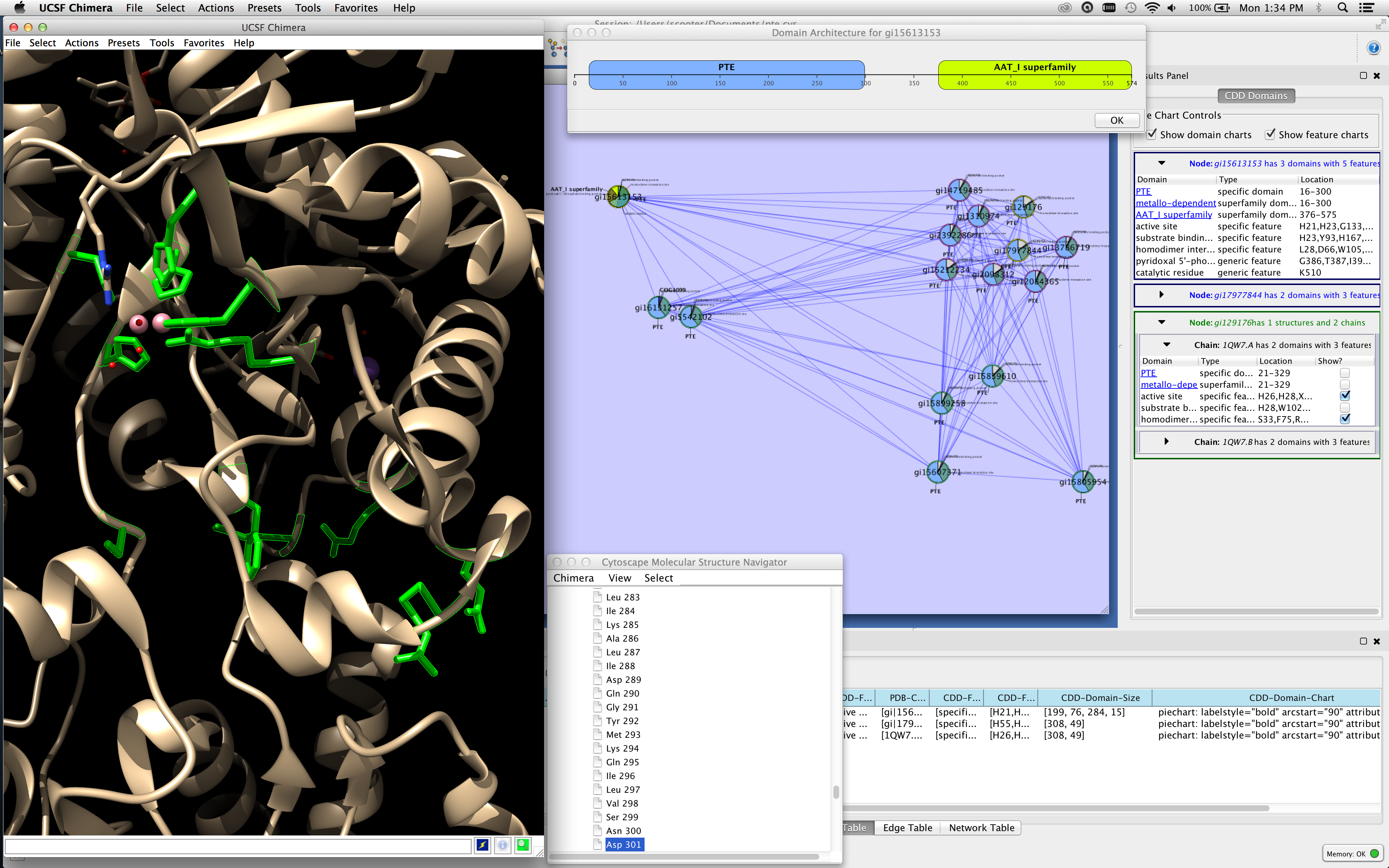
User-defined interaction types are also supported. For yeast and other model organisms, large sources of pairwise interactions are available through the BIND and TRANSFAC databases.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
